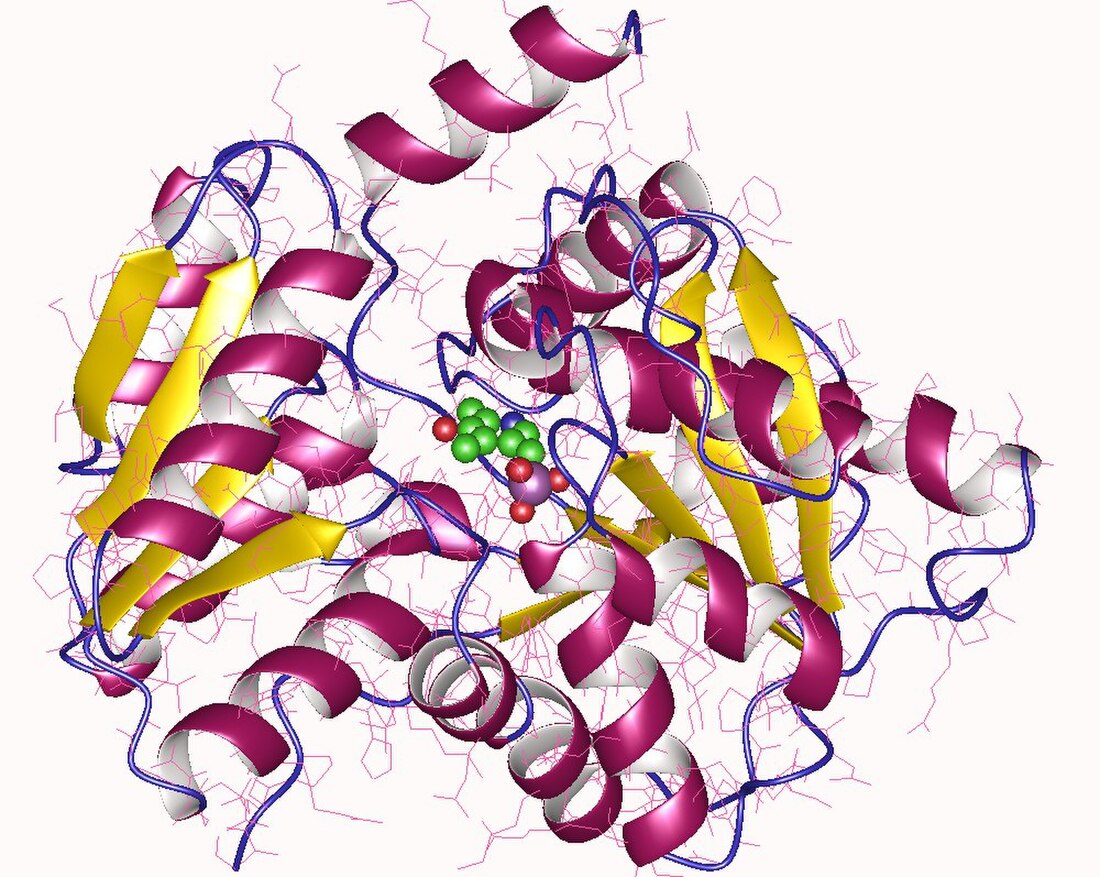
The enzyme L-serine ammonia-lyase (EC 4.3.1.17) catalyzes the chemical reaction
- L-serine = pyruvate + NH3 (overall reaction)
- (1a) L-serine = 2-aminoprop-2-enoate + H2O
- (1b) 2-aminoprop-2-enoate = 2-iminopropanoate (spontaneous)
- (1c) 2-iminopropanoate + H2O = pyruvate + NH3 (spontaneous)
| L-serine ammonia-lyase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Serine dehydratase monomer, Human | |||||||||
| Identifiers | |||||||||
| EC no. | 4.3.1.17 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
This enzyme belongs to the family of lyases, specifically ammonia lyases, which cleave carbon-nitrogen bonds. The systematic name of this enzyme class is L-serine ammonia-lyase (pyruvate-forming). Other names in common use include serine deaminase, L-hydroxyaminoacid dehydratase, L-serine deaminase, L-serine dehydratase, and L-serine hydro-lyase (deaminating). This enzyme participates in glycine, serine, threonine and cysteine metabolism. It employs one cofactor, pyridoxal phosphate.
Structural studies
As of late 2007, 4 structures have been solved for this class of enzymes, with PDB accession codes 1P5J, 1PWE, 1PWH, and 2IQQ.
References
Wikiwand in your browser!
Seamless Wikipedia browsing. On steroids.
Every time you click a link to Wikipedia, Wiktionary or Wikiquote in your browser's search results, it will show the modern Wikiwand interface.
Wikiwand extension is a five stars, simple, with minimum permission required to keep your browsing private, safe and transparent.